The aim of the study was to identify the isolation rate of thermotolerant campylobacters in a small-scale broiler-meat production farm over a one-year period. The second deliverable of the study was to determine the potential virulence markers. The laboratory investigation was performed on 283 samples (cloacal swabs, caeca, carcass swabs) collected on three sampling points (farm, slaughter line, and cold storage). The isolates obtained with the conventional microbiological method were confirmed with multiplex PCR for identification of campylobacters. The presence of 10 virulence genes was analyzed in the C. jejuni isolates ( flaA, racR, virB11, dnaJ, wlaN, cadF, ciaB, cdtA, cdtB, cdtC). Out of 283 samples, 169 (59.7%) were confirmed as Campylobacter spp., 111 (39.2%) C. jejuni, and 43 (15.2%) C. coli. C. jejuni was the most prevalent in all sampling points. Campylobacter spp. showed a characteristically seasonal prevalence with the highest isolation rate during the warmer period of the year. We detected the cadF and ciaB genes in all C. jejuni isolates. The flaA gene was present in 50% of the examined strains. The cdt genes (cdtA, cdtB, and cdtC) were confirmed in 52.8%, 52.8%, and 47.2% of the C. jejuni strains, respectively. C. jejuni showed 15 profiles of virulence patterns with four predominant profiles.
Campylobacter is the most commonly reported causative agent for human bacterial diarrhoeal disease in the European Union since 2005 (
5). The main source of infection is by consumption of food with animal origin, especially undercooked poultry meat, unpasteurized milk, or cross-contaminated meat products during preparation and storage (
6). Besides these risk factors there are several more that are worth considering: abroad traveling, exposure to animals, working in slaughterhouses, animal farms visiting, consumption of pork meat, contact with dogs and cats, and swimming in open waters (
7, 8, 9, 10).
The infection is usually brief, mild, self-limiting, and generally does not require any treatment. In some cases, the infection is associated with serious clinical manifestations such as bacteremia, reactive arthritis, hemolytic uremic syndrome, meningitis, septicemia, and Guillain-Barré syndrome (
1, 11). Macrolides and fluoroquinolones are frequently used as the first and second-choice drugs for highrisk patients (young, elderly, immunocompromised individuals) (
12).
Many authors have confirmed the seasonal pattern variation in the occurrence rate of
Campylobacter. The usual prevalence spikes take place during the warmer months of the year and are generally associated with the higher survival of
Campylobacter in the habitat (
13).
The molecular pathogenic mechanism in
Campylobacter infection is still not perfectly understood (
14). The extensive research on campylobacters has revealed numerous genes relevant for their virulence and pathogenicity (
15):
flaA, flhA, cadF, dnaJ, and
racR (genes that are engaged in the stage of adherence and colonization);
virB11,
ciaB, and
iam (genes important for the invasiveness);
cdtA, cdtB, and
cdtC (genes bound to the cytotoxic effect on the epithelial cells), and
wlaN (linked with the synthesis of the lipooligosaccharides, resulting in Guillain-Barré Syndrome) (
16).
The objectives of this study were to 1) estimate the isolation rate of thermotolerant
Campylobacter in broilers sampled in farms and slaughterhouse; 2) identify (if present) the seasonal variation in
Campylobacter prevalence; 3) detect the presence of virulence-related genes in
Campylobacter jejuni isolates, and 4) evaluate the detected virulence patterns in
Campylobacter jejuni isolates.
MATERIAL AND METHODS
Collection of samplesThis study was conducted on one broiler farm and slaughterhouse in the Skopje region, N. Macedonia, in the period between March and December 2017. All procedures involving animals were in accordance with the ethical standards and international guidelines for the care and use of animals.
A total of 283 samples were collected. The farm had a capacity of 10.000 broilers with a conventional system. The usual rate of slaughtering was by several thinning cycles of 1.500 broilers per day.
Cloacal swabs were taken from randomly chosen broilers at the farm one week prior to slaughter (n=64). The samples were placed in plastic tubes containing 5 mL of Preston broth.
Broiler caeca were collected at the slaughter line (n=166) during the evisceration phase, and were packed in sterile plastic bags. Samples were collected randomly from at least 40 birds in each sample-collection cycle during the slaughtering process.
Swab samples from broiler carcasses (n=53) were taken in the storage area before shipping to the consumers (cold chamber of the slaughterhouse) using sterile cotton swabs, which were placed in plastic tubes containing 5 mL of Preston broth. The samples were collected the following day after slaughtering, including at least 10 carcasses in each sample-collection cycle.
Samples were stored and transported to the laboratory at 4-8 °C. The laboratory analysis was initiated 3-4 hours after sampling. Isolation and confirmation of Campylobacter spp.
The isolation and identification of thermotolerant Campylobacter was performed according to the standard ISO 10272-1:2017 (
17). The positive isolates were sub-cultured on mCCDA agar plates and stored in glycerol broth at -80 °C.
Extraction of DNA and PCR analysisThe DNA preparation for the
Campylobacter isolates was performed by the boiling method. The procedure included suspension of the cultures in 0.5 mL of TE buffer, boiling at 95 °C for 10 min, and centrifugation at 15.000 rpm for 5 minutes. Acquired supernatants were kept at -20 °C and used for both PCR methods.
The multiplex PCR for identification and differentiation of
Campylobacter spp. was performed with adaptations according to the previously published study (
18). This multiplex PCR could identify several
Campylobacter species (
C. jejuni subsp. jejuni;
C. coli, C. lari, C. upsaliensis, and
C. fetus subsp. fetus). The list of the used primers is shown in
Table 1.
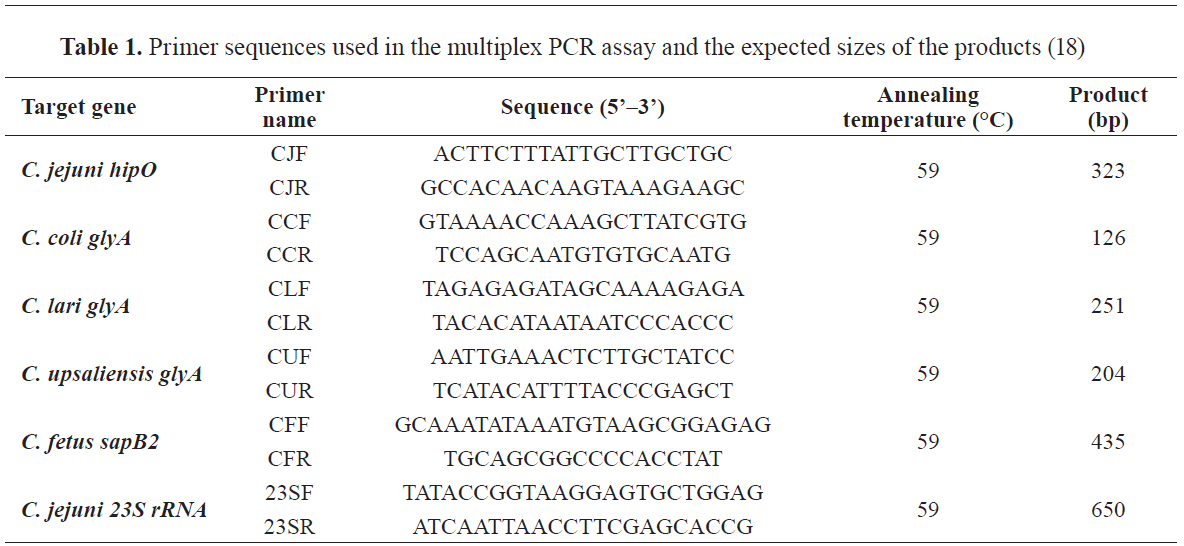

PCR amplifications were performed in a mixture (25 μL) consisting of 12.5 μL of 2× Platinum Multiplex PCR Master Mix (Applied Biosystems, UK), 2.5 μL of template DNA, and 5 μL of primer mix (0.5 μM
C. jejuni and
C. lari primers; 1 μM
C. coli and
C. fetus primers, 2 μM
C. upsaliensis primers; 0.25 μM 23S rRNA primer). Distilled water was added to make 25 μL. DNA amplification was carried out in a thermocycler (Techne, UK) using an initial denaturation step at 95 °C for 2 min followed by 35 cycles of amplification (denaturation at 95 °C for 30 sec., annealing at 59 °C for 45 sec., and extension at 72 °C for 45 sec.), ending with a final extension at 72 °C for 10 min.
The multiplex PCR for detection of the virulence genes in the
C. jejuni isolates was performed as proposed by Datta (19). The list of the used primers is shown in
Table 2.
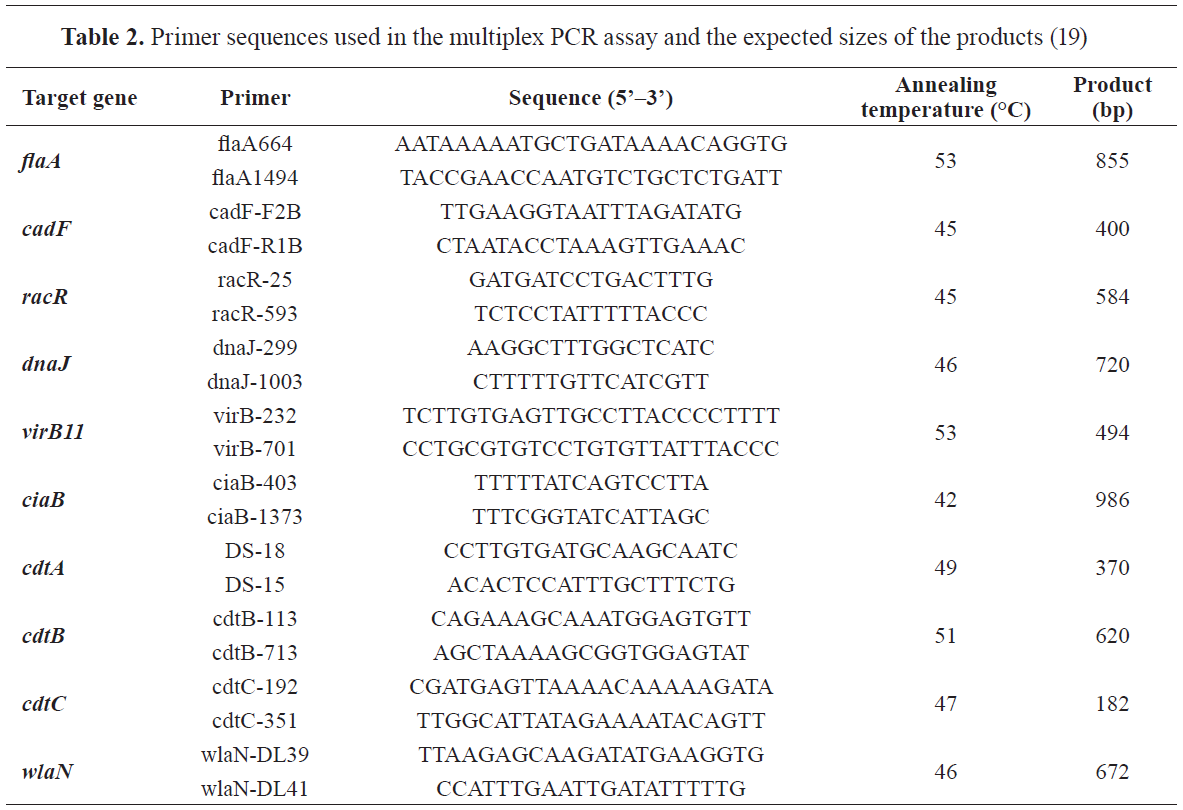
For this protocol each multiplex PCR tube contained 25 μL of mixture: 12.5 μL Platinum Multiplex PCR Master Mix (Applied Biosystems, UK), 2.5 μL of template DNA, 5 μL of primer mix (0.5 μM of the used primers), and 5 μL of distilled water. Amplification was performed with initial denaturation step at 95 °C for 2 min followed by 35 cycles of amplification (denaturation at 94 °C for 1 min, annealing at a temperature specific to the primer pair for 1 min, and extension at 72 °C for 1 min) and final extension at 72 °C for 10 min.
RESULTS
Isolation rate of thermotolerant Campylobacter, C. jejuni, and C. coli at broiler farm and at the slaughterhouseThe results of the study indicated that
Campylobacter spp. was present in all phases of the production. The distribution of campylobacters was highest in the cloacal swabs (73.4%). The results of the laboratory examinations showed that
C. jejuni was the most prevalent species on the farm and in the slaughterhouse (
Table 3).
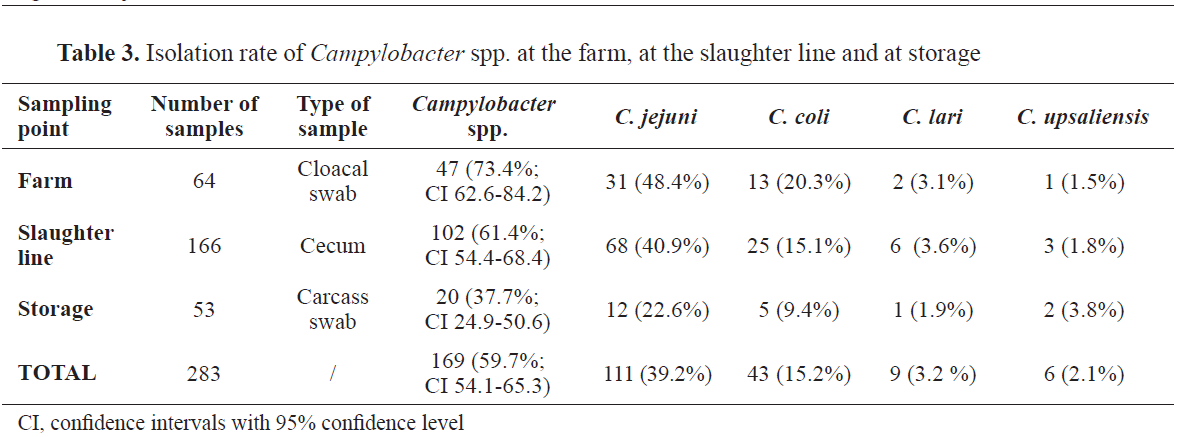
This study identified high prevalence of
Campylobacter in carcasses (37.7%), consisting mainly of isolates of
C. jejuni (22.6%), followed by
C. coli (9.4%).
Seasonal variation in Campylobacter spp. prevalenceAs previously mentioned, in several studies,
Campylobacter spp. has shown a seasonal trend of higher prevalence in the warmer period of the year. In the current study (
Fig. 1) we confirmed this trend with highest prevalence in the samples taken in August (75.3%), followed by the June sampling period (67.1%).
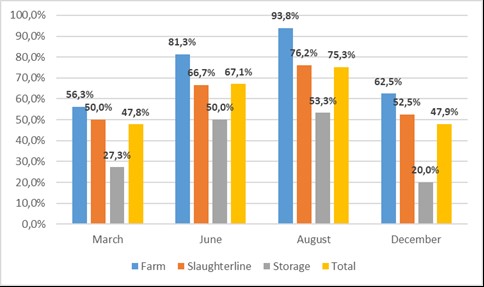 Figure 1.
Figure 1. Seasonal prevalence of
Campylobacter spp. along the broiler chain
Presence of virulence genes in C. jejuniAmong the isolates, 111 were confirmed as
C. jejuni by both the classical method and the multiplex PCR. Three of the isolates were dismissed because of technical issues with the template DNA purity. Therefore, 108 isolates were subjected to further analysis for the detection of virulence genes (
Fig. 2;
Table 4).
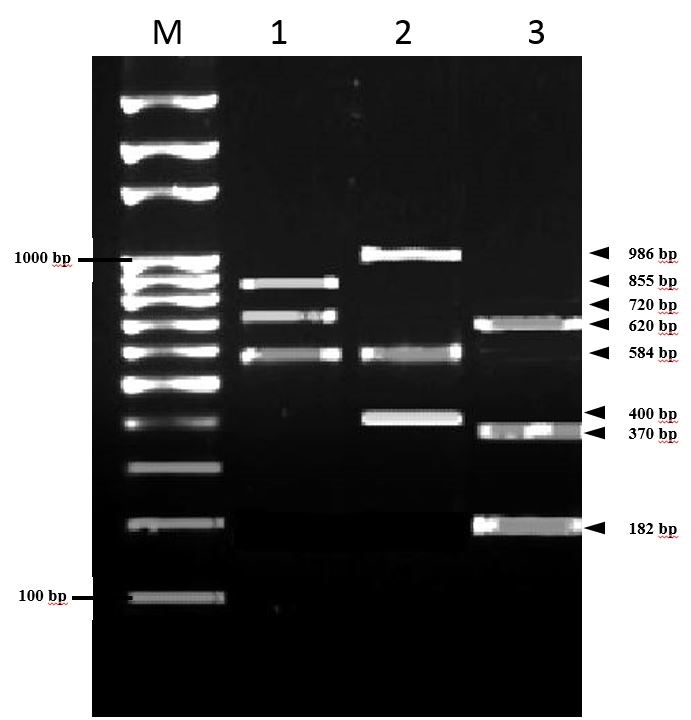 Figure 2.
Figure 2. Visualization of PCR amplicons of virulence genes. Lane marked with M: 100-bp ladder;
Lane 1:
racR, dnaJ and
flaA genes; Lane 2:
cadF, racR and
ciaB genes; Lane 3:
cdtC, cdtA and
cdtB genes
 Evaluation of the detected virulence patterns in C. jejuni
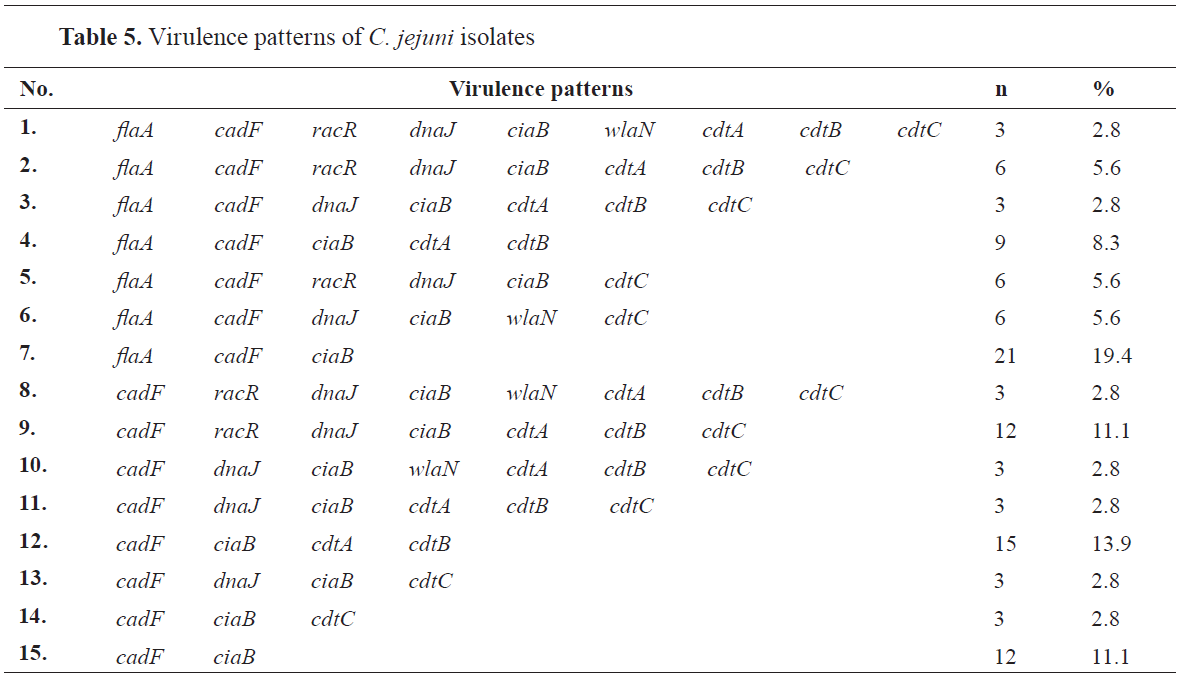
Evaluation of the detected virulence patterns in C. jejuniIn total, we detected 15 virulence patterns in the
C. jejuni isolates. In 19.4% of the confirmed
C. jejuni isolates, the analysis detected the profile No. 7. The second highest prevalence was detected for the profile No. 12 (in 13.9% of the isolates). The profile No. 1 (all of the virulence genes present except
virB11) was confirmed only in 3 isolates (2.8%) of
C. jejuni (
Table 5).
DISCUSSION
In this study,
Campylobacter spp. was present in all type of samples collected along the broiler meat production chain (cloacal swabs, caeca, and carcass swabs). The confirmed level of
Campylobacter spp. on the broiler farm was very similar with the level confirmed in two studies performed on farms with comparable breeding conditions (
20, 21).
This study also revealed high isolation rate of
Campylobacter spp. (61.4%) in the slaughterhouse (evisceration phase) in the cecum samples. Analysis of cecal samples showed a great variation in the isolation rate (region, age of the flock, seasonality, on farm pre-harvest measures, farm management) (
22). In some studies, the isolation rate was in accordance with our results (
23), but there are studies that report even higher isolation rate (
24).
At the last point of sampling (cold storage), 37.7% of the samples were positive. The point of concern is that 22.2% of the samples have shown to be positive for
C. jejuni. This is important as from this point the poultry meat is dispatched to the consumers. If we follow the
Campylobacter spp. presence along the production chain, we find reduced level but not eliminated level of contamination (
25).
Our results have confirmed a well-defined seasonal pattern of flock colonization with
Campylobacter spp. As observed by many authors, it is characterized with highest prevalence in summer or autumn (
26, 27, 28). The mechanism by which temperature affects
Campylobacter colonization of broilers is probably linked with microbial survival, higher numbers of wildlife vectors, increased ventilation and fan speeds, insects, and rodents (
29).
The pathogenicity of thermotolerant
Campylobacter found in the broiler meat in addition to its isolation rate gives a clear picture of the public health risk. In this study we detected several of the virulence genes in
C. jejuni isolates. Our results showed that
cadF and
ciaB genes were detected in all of the confirmed strains (
Table 5). Similar observations indicated that the
cadF and
ciaB genes were present in the
Campylobacter isolates from chicken carcasses and droppings (
19, 30).
The analyses detected an isolation rate of 50% of the
flaA gene which coordinate the motility and virulence (
31, 32). Comparable levels of prevalence of this gene were detected in other studies (
33, 34).
The frequency of
cdt genes (52.8%, 52.8%, and 47.2%) that was observed in our
C. jejuni isolates was close to the prevalence detected in studies by other authors (
35, 36). This holotoxin (cdtABC) produced by campylobacters is a factor which has a great role in the development of the disease. All three toxin subunit genes (
cdtA, cdtB, and
cdtC) are necessary for the cytotoxin expression (
37).
The
wlaN gene (prevalence of 13.9%) is responsible for the metabolism of specific lipooligosaccharides (LOS) connected with disease sequelae such as Guillain-Barre syndrome (
38, 39). Concerning the genes connected with adherence and colonization (
40) the study results confirmed 44.4% prevalence of the
dnaJ gene and lower prevalence of the
racR gene (27.8%). The
virB11 gene, which is responsible for the expression of invasion (
41), was not detected in the
C. jejuni isolates.
The authors acknowledge that at the time of sample collection for the purpose of the current research, the country had a small-scale broiler production based on few farms and only one operating slaughterhouse imposing limitations of sample number.
CONCLUSION
This study confirmed high isolation rate of
Campylobacter spp. in the broiler meat production chain. This statement is supported both at the farm and the slaughterhouse (evisceration phase and cold storage) in different type of analyzed samples.
C. jejuni was more frequently isolated than
C. coli at the three sampling points.
The
Campylobacter isolation rate in the cold chamber samples was notable and relevant for public health consideration.
C. jejuni isolates had varying virulence patterns (n=15) and diverse pathogenic potential which are not always necessarily expressed in-vivo. The detected virulence genes and their patterns surely demonstrate the pathogenic potential of
Campylobacter jejuni isolated across the broiler chain.
CONFLICT OF INTEREST
The authors declare that they have no potential conflict of interest with respect to the authorship and/or publication of this article.
ACKNOWLEDGMENTS
This research was supported by the Faculty of Veterinary Medicine – Skopje. The authors would like to express their gratitude to colleagues Prof. Kiril Krstevski and Prof. Igor Dzadzovski for their scientific advice and technical help.
AUTHORS’ CONTRIBUTIONS
LjA conceived and designed the study and wrote the manuscript. LjA, ZP, KB and MP performed the experiments. SM contributed to the final version. DJ gave critical revision. PS supervised the project and approved it for publishing.

 10.2478/macvetrev-2021-0020
10.2478/macvetrev-2021-0020






