Original Scientific Article
Molecular characterization of the emerging sublineage Ind2001e of Foot-and-mouth disease virus serotype O in buffalo in Manikganj, Bangladesh
Md Zulfekar Ali
*
 ,
,
Naima Tasnim
 ,
,
Md Ataur Rahman
 ,
,
Md Hafizur Rahman
 ,
,
Mirza Mienur Meher
 ,
,
Md Giasuddin

Received: 13 December 2024
Received in revised form: 27 April 2025
Accepted: 30 July 2025
Available Online First: 25 August 2025
Published on: 15 October 2025
Correspondence: Md Zulfekar Ali, zulfekarvet@gmail.com
Abstract
Foot-and-mouth disease (FMD) is a major economic infectious disease in endemic regions like Bangladesh, affecting cloven-hooved animals and hindering livestock development. Good farm practices, including vaccination, is the only effective control method and continuous monitoring is necessary to detect vaccine effectiveness. Buffalo, a significant livestock resource in the southern coastal region of Bangladesh, requires more disease management efforts than cattle. This study aimed to isolate and molecularly characterize the VP1 coding region of FMDV in clinically suspected buffaloes. Tongue epithelial tissues were collected from two buffaloes suspected of FMD in June-2018 in the Manikganj district of Bangladesh. Both samples were identified as FMDV serotype O using RT-PCR testing and were subsequently cultured in BHK-21 cell line. Phylogenetic analysis of VP1 region revealed that both isolates clustered into a distinct sublineage Ind2001e, within the dominant lineage Ind2001 of Middle East-South Asia (ME-SA) topotype of FMDV serotype O, which has been circulating in Bangladesh since 2012. Analysis of nucleotide (NT) similarities revealed that 92.38% of NTs are homologous between isolates, whereas similarities with the Indian sequences were 99.91%-99.99%, and with the Nepal sequences it was 99.90%-99.99%. The amino acid mutations within the VP1 sequence between the vaccine strain (O/India/R2/75) and the BAN450 isolates was 3.75% (8; 96.25% homology), specifically in the BC loop and C-terminal region. In conclusion, buffaloes get infected with the emerging novel sublineage Ind2001e. The effectiveness of vaccination in Bangladesh with the potential mutational trend poses significant concern regarding the transmission between multiple species.
Keywords: buffalo, Foot-and-mouth-disease, Ind2001e sublineage, molecular characterization, serotype O
INTRODUCTION Foot-and-mouth disease (FMD) is a transboundary, airborne, acute, and highly contagious animal disease characterized by high fever and anorexia, followed by vesicular lesions in the oral mucosa and interdigital spaces of cloven-hoofed animals, such as buffalo, cattle, goats, pigs, sheep, and various wild species ( 1 , 2 ). Consequently, the animal undergoes weight loss, resulting in reduced production, particularly in milk and meat production. FMD can also cause abortion, death in calves, international trade restrictions, and high treatment costs ( 2 , 3 ). It has a significant global economic impact which is estimated between 6.5 and 21 billion US dollars. In endemic countries it can reach higher than 1.5 billion dollars annually ( 4 ). A study by Giasuddin et al. ( 5 ) projected an annual economic impact of 2.22 billion dollars in Bangladesh. Foot-and-mouth disease virus (FMDV) is the etiological agent comprising seven serotypes characterized by distinct genetic and antigenic variations: O, A, Asia 1, C, SAT-1, SAT-2, and SAT-3. Currently, only serotypes O, A, and Asia 1 are prevalent in Bangladesh, with serotype O predominating in 82% of the outbreaks ( 6 ). Serotype O is divided into 11 topotypes of which the Middle-East-South Asia (ME-SA) topotype was identified in Bangladesh. To date, four lineages within the ME-SA topotype have been identified: Ind2001, Ind2011, PanAsia, and PanAsia-2 ( 7 , 8 ). The Ind2001 lineage subsequently developed into five sublineages: Ind2001a, b, c, d, and e (9, 10). A newly identified sublineage Ind2001BD1, was reported to be the predominant sublineage in Bangladesh which was later classified as Ind2001e by the World Reference Laboratory for Foot-and-mouth disease (WRLFMA) ( 11 , 12 ). However, to prevent FMDV in cattle and buffaloes, the Government of Bangladesh has adopted a national strategic plan to control FMD through early diagnosis, strategic mass vaccination, zonal control measures, and implementation of the OIE FMD control model. Although most of the research focuses on FMDV in cattle, there is less synthesized information on monitoring of FMDV in buffaloes, despite being the second largest livestock population in Bangladesh. Buffalo, referred to as "Black Gold," holds significant value and is predominantly located in the southern coast area of Bangladesh ( 13 , 14). The buffalo management system remains reliant on traditional practices and lacks modern housing, biosecurity measures, nutritional management, and disease control strategies. Farmers have misconceptions regarding the necessity of vaccinations and deworming for buffaloes. Consequently, buffaloes were silently affected by many infectious diseases, including FMD, yet remained underreported ( 15 ). Therefore, the study aimed to isolate and molecularly type serotypes, and characterize FMDV serotype O, including amino acid mutations in the VP1 coding region, in comparison to the vaccine strain identified in buffaloes from the Manikganj district of Bangladesh. Based on the previous reports of genetic diversity of FMDV circulating in Bangladesh, we hypothesized that the FMDV serotype isolated from buffaloes in Manikgonj will carry notable mutations in the VP1 coding region which may reduce the match with the vaccine strain. MATERIAL AND METHODS
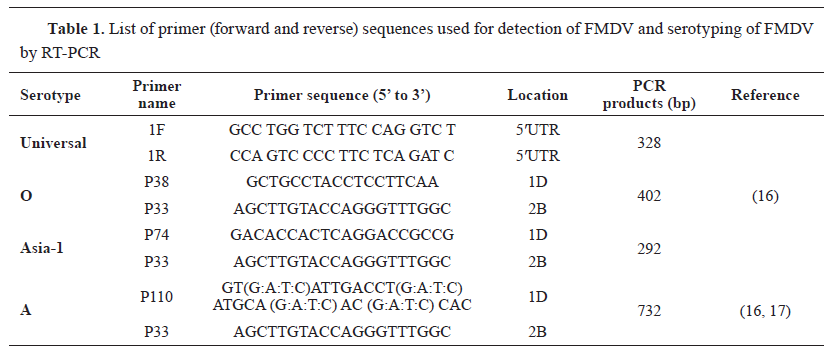
Sample collection Two tongue epithelial tissue samples were aseptically collected into virus transfer medium from two different buffalo farms suspected of having FMD in June 2018, located in the Manikganj District (coordinates: 23.85°N 90.01°E) of Bangladesh ( Fig. 1 ). Clinical signs in buffaloes include absence of appetite, fever, drooling of saliva, vesicles on the tongue and gums. The animals were not vaccinated and were mixed with cattle that have recently recovered from FMD. Samples were immediately transferred to the laboratory in a cooler with ice and stored at -80 °C in a freezer until subsequent laboratory procedures. Detection of FMDV serotypes by RT-PCR test The tongue epithelial samples were ground and macerated in a pestle and mortar with sterile sand. A 10% tissue suspension was made with phosphate-buffered saline. Total RNA was extracted from an aliquot of suspension samples using the QIAamp® Viral RNA kit (QIAGEN, Hilden, Germany) following the manufacturer's instructions. The VP1 gene-based PCR diagnostic assay was used for the confirmation of FMDV, adopting the primers and RT-PCR protocols developed by Vangrysperre and De Clercq (16), with minor modifications including the QIAGEN® One-Step RT-PCR Kit (QIAGEN, Hilden, Germany) as the RT-PCR master mix. Additionally, three serotypes A, O, and Asia 1 of FMDV circulating in Bangladesh were examined using the primers and procedures designed by Vangrysperre and De Clercq ( 16 ) and Callens and De Clercq ( 17 ). The list of primers is described in Table 1 , and the RT-PCR temperature profile is described in Table 2.
Isolation of FMDVThe tissue inoculums were inoculated onto a pre-established monolayer of BHK-21 cells (widely used cell line for FMDV culture) using Eagle's medium supplemented with trypticase soy broth and antibiotics, incubated at 37 °C in a 5% CO 2 condition with 10% fetal bovine serum (Gibco, USA). The cells were monitored consistently until the third passage, and after four days post-inoculation. Characteristic cytopathic effects (CPE) became apparent, including cell rounding and flattening, formation of multinucleated giant cells, disruption of intracellular connections, and cell death ( 18 ).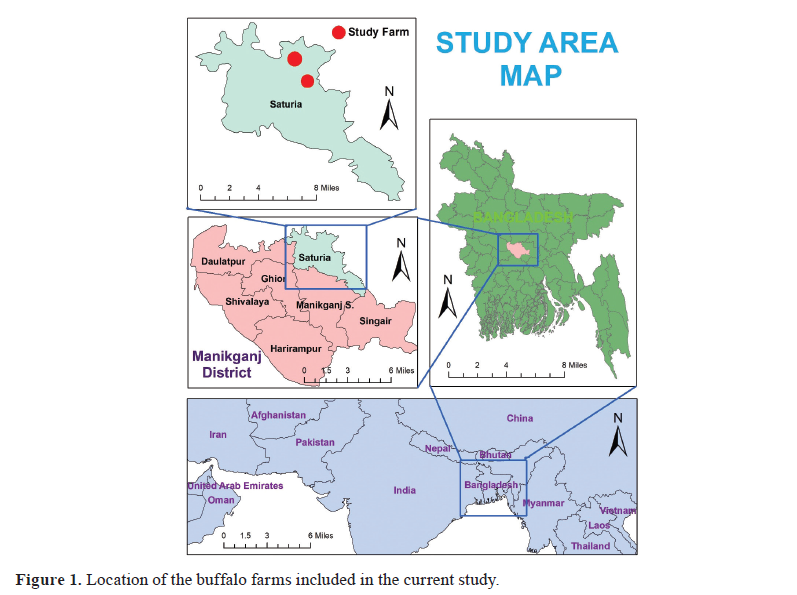
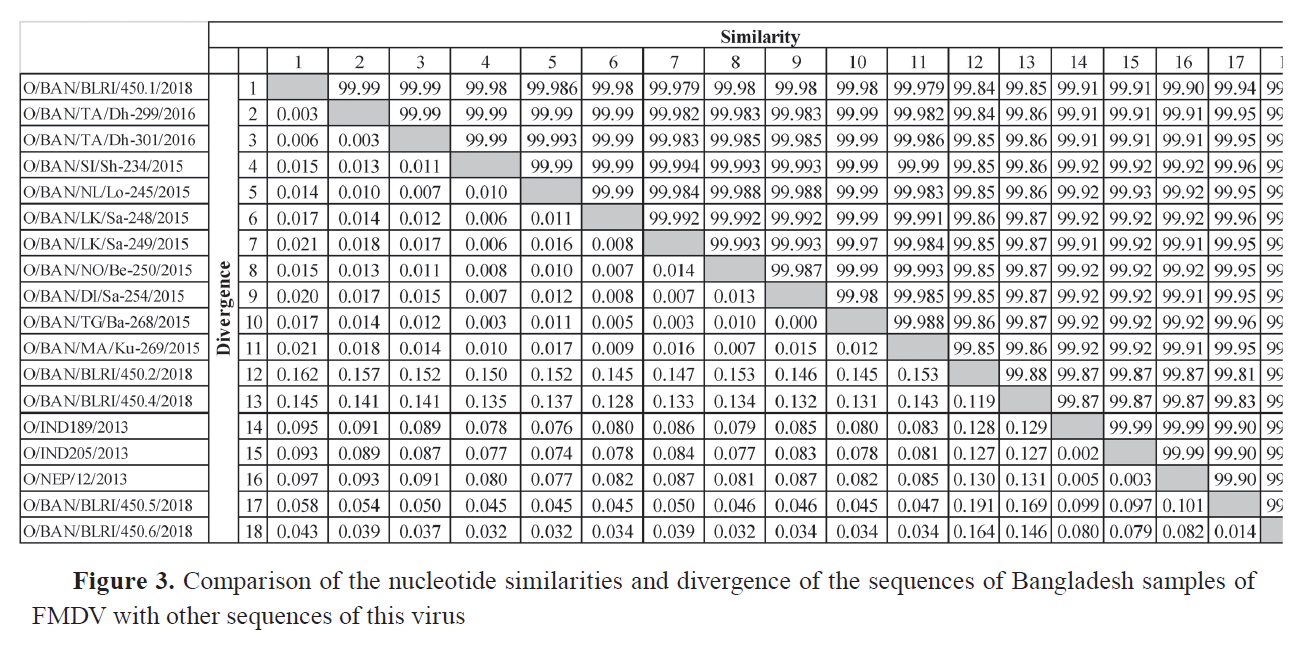
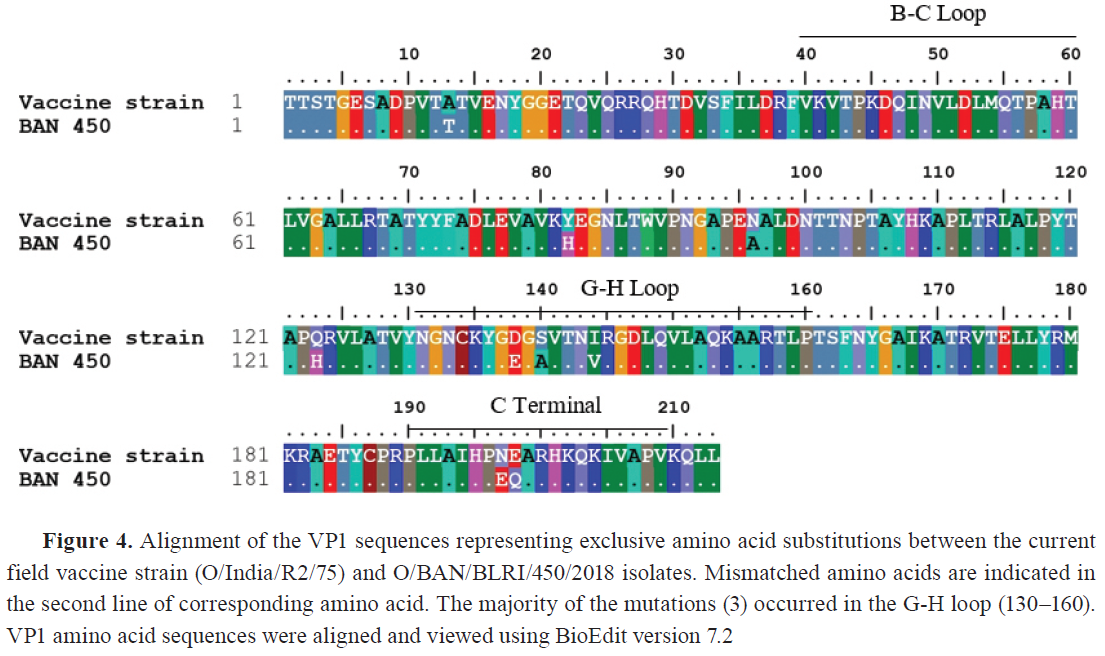
 Sequencing of VP1 coding region Total RNA was extracted from the third passages of both isolates using the QIAamp® Viral RNA kit. The VP1 gene coding region of FMDV was amplified using three sets of primers and PCR methods as described by Knowles et al. ( 19 ) for FMDV serotype O. The amplified PCR products were analyzed using electrophoresis on a 1.5% agarose gel, alongside a 1kb DNA ladder stained with SYBR Safe DNA Gel Stain (ThermoFisher, Waltham, MA), and a UV transilluminator confirmed the amplified band. Later on, the QIAquick® Gel Extraction Kit (QIAGEN®, Hilden, Germany) was used to purify the amplified products. The products were quantified and sent to the commercial source for cycle sequencing reaction of amplified FMD serotype O products. The forward and reverse sequences of the VP1 coding region were subjected to quality assessment, editing, and trimming. They were assembled into a consensus sequence by BioEdit version 7.2. Phylogeny analysis A total of 56 VP1 coding sequences were retrieved from the GenBank database through NCBI BLAST ( 20 ) search, sourced from neighboring countries and Bangladesh, for an analysis of the evolutionary divergence of FMDV serotype O isolated in this study. The sequences were assembled and aligned using the ClustalW algorithm ( 21 ) using MEGA11 software ( 22 ). Finally, the phylogenetic tree with different lineages and sublineages was constructed using MEGA11. The best substitution model Tamura-Nei ( 23 ), with discrete Gamma distribution (TN93+G) in the Maximum Likelihood method, with 1,000 times bootstrap replications, was applied based on nucleotide alignment data. Mutational analysis The VP1 coding amino acid sequence of the BAN 450 (O/BAN/BLRI/450/2018) isolate was compared with the common vaccine strain O/India/R2/75 for checking the mutational spectrum between them by using BioEdit 7.2. The amino acid sequence was translated from the nucleotide sequence based on the standard genetic code after codon-based alignment using MEGA11 software ( 22 ). Ethical approval The research protocol was approved by the Animal Experimentation Ethics Committee, Bangladesh Livestock Research Institute, Savar, Dhaka 1341, Bangladesh, under reference-BLRI0006. RESULTS The VP1-based RT-PCR test confirmed that the two tongue epithelium samples obtained from both buffalo farms were positive for FMDV. Subsequent serotyping via RT-PCR revealed that both were confirmed as FMDV serotype O. The VP1 coding sequences of positive isolates were submitted to the NCBI GenBank database with accession IDs PQ468693 (O/BAN/BLRI/450.5/2018) and PQ468694 (O/BAN/BLRI/450.6/2018) (October 2024). The VP1 sequence data analysis for FMDV using the NCBI BLAST database revealed 100% nucleotide query coverage with FMDV serotype O. The 26 sequences of FMDV serotype O, including the two sequences of the current study, were clustered within the emerging novel sublineage Ind2001e from lineage Ind2001 under the Middle East-South Asia (ME-SA) topotype ( Fig. 2 ). Phylogenetic analysis showed that the isolates identified in this study clustered within the same group, corresponding to FMDV serotype O strains previously reported in Bangladesh between 2012 and 2018. The VP1 coding region sequences had a nucleotide similarity of 92.38%. The VP1 coding region sequences of FMDV serotype O obtained from NCBI GenBank had nucleotide similarity of 99.10% to 99.97% with various Bangladeshi isolates, 99.91% to 99.99% with Indian isolates, and 99.90% to 99.99% with Nepalese sequences ( Fig. 3 ). Eight mutations were identified among the isolates and the vaccine strains BAN 450 and O/India/R2/75, with the majority occurring in the GH loop (D138E, S140A, S140H, I144V) in the VP1 regions of buffalo isolates. Two mutations were seen in the C-terminus (N197E and E198Q), while no mutations were found in the BC loop ( Fig. 4 ). Cytopathic effects were observed in the cell line ( Fig. 5 ), and the presence of the virus was subsequently confirmed by RT-PCR.
Sequencing of VP1 coding region Total RNA was extracted from the third passages of both isolates using the QIAamp® Viral RNA kit. The VP1 gene coding region of FMDV was amplified using three sets of primers and PCR methods as described by Knowles et al. ( 19 ) for FMDV serotype O. The amplified PCR products were analyzed using electrophoresis on a 1.5% agarose gel, alongside a 1kb DNA ladder stained with SYBR Safe DNA Gel Stain (ThermoFisher, Waltham, MA), and a UV transilluminator confirmed the amplified band. Later on, the QIAquick® Gel Extraction Kit (QIAGEN®, Hilden, Germany) was used to purify the amplified products. The products were quantified and sent to the commercial source for cycle sequencing reaction of amplified FMD serotype O products. The forward and reverse sequences of the VP1 coding region were subjected to quality assessment, editing, and trimming. They were assembled into a consensus sequence by BioEdit version 7.2. Phylogeny analysis A total of 56 VP1 coding sequences were retrieved from the GenBank database through NCBI BLAST ( 20 ) search, sourced from neighboring countries and Bangladesh, for an analysis of the evolutionary divergence of FMDV serotype O isolated in this study. The sequences were assembled and aligned using the ClustalW algorithm ( 21 ) using MEGA11 software ( 22 ). Finally, the phylogenetic tree with different lineages and sublineages was constructed using MEGA11. The best substitution model Tamura-Nei ( 23 ), with discrete Gamma distribution (TN93+G) in the Maximum Likelihood method, with 1,000 times bootstrap replications, was applied based on nucleotide alignment data. Mutational analysis The VP1 coding amino acid sequence of the BAN 450 (O/BAN/BLRI/450/2018) isolate was compared with the common vaccine strain O/India/R2/75 for checking the mutational spectrum between them by using BioEdit 7.2. The amino acid sequence was translated from the nucleotide sequence based on the standard genetic code after codon-based alignment using MEGA11 software ( 22 ). Ethical approval The research protocol was approved by the Animal Experimentation Ethics Committee, Bangladesh Livestock Research Institute, Savar, Dhaka 1341, Bangladesh, under reference-BLRI0006. RESULTS The VP1-based RT-PCR test confirmed that the two tongue epithelium samples obtained from both buffalo farms were positive for FMDV. Subsequent serotyping via RT-PCR revealed that both were confirmed as FMDV serotype O. The VP1 coding sequences of positive isolates were submitted to the NCBI GenBank database with accession IDs PQ468693 (O/BAN/BLRI/450.5/2018) and PQ468694 (O/BAN/BLRI/450.6/2018) (October 2024). The VP1 sequence data analysis for FMDV using the NCBI BLAST database revealed 100% nucleotide query coverage with FMDV serotype O. The 26 sequences of FMDV serotype O, including the two sequences of the current study, were clustered within the emerging novel sublineage Ind2001e from lineage Ind2001 under the Middle East-South Asia (ME-SA) topotype ( Fig. 2 ). Phylogenetic analysis showed that the isolates identified in this study clustered within the same group, corresponding to FMDV serotype O strains previously reported in Bangladesh between 2012 and 2018. The VP1 coding region sequences had a nucleotide similarity of 92.38%. The VP1 coding region sequences of FMDV serotype O obtained from NCBI GenBank had nucleotide similarity of 99.10% to 99.97% with various Bangladeshi isolates, 99.91% to 99.99% with Indian isolates, and 99.90% to 99.99% with Nepalese sequences ( Fig. 3 ). Eight mutations were identified among the isolates and the vaccine strains BAN 450 and O/India/R2/75, with the majority occurring in the GH loop (D138E, S140A, S140H, I144V) in the VP1 regions of buffalo isolates. Two mutations were seen in the C-terminus (N197E and E198Q), while no mutations were found in the BC loop ( Fig. 4 ). Cytopathic effects were observed in the cell line ( Fig. 5 ), and the presence of the virus was subsequently confirmed by RT-PCR.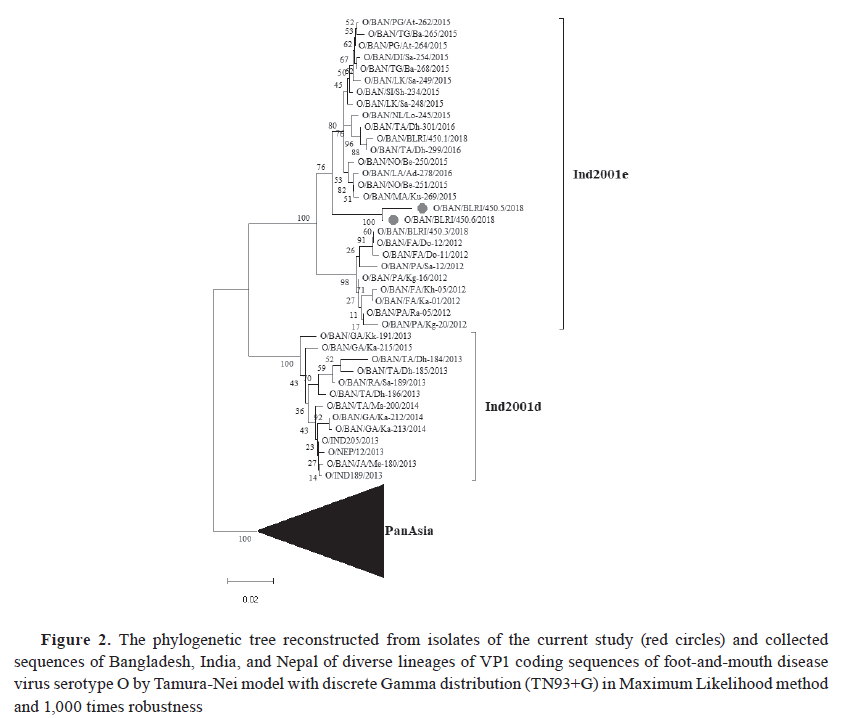
 DISCUSSION FMD affects all cloven hoof animal species in Bangladesh every year, especially cattle and buffalo. It has been reported that the genetic evolution of FMDV leads to a consistently increasing tendency of infection and mortality ( 24 , 25 ). Buffalo were perceived as having a lower susceptibility to diseases, and farmers considered buffaloes resistant to FMDV ( 26 ). Evidence indicates that Bangladeshi buffalo farmers occasionally experience outbreaks of FMD, although symptoms are less severe than in cattle ( 6 ). In recent outbreaks, FMDV serotype O has been the predominant serotype in both cattle and buffalo in Bangladesh ( 12 ). However, there is a lack of comprehensive genetic information on FMDV serotype O in buffalo as compared to cattle in Bangladesh. The study aimed to investigate the evolutionary relationships of FMDV serotype O circulating in buffalo in Bangladesh and to compare the amino acid mutation against the vaccine strain based on the VP1 coding sequence of the genome. The VP1 sequencing analysis indicated that the lineage Ind2001 has been predominantly circulating in Bangladeshi cattle since 2009 ( 9 , 25 ). The study identified an emerging novel sublineage Ind2001e (previous name Ind2001BD1) under the ME-SA topotype circulating in both buffalo and cattle. The sublineages Ind2001e and Ind2001BD2 were first identified in samples collected between 2012 and 2016 by Siddique et al. ( 9 ). Later, lineage PanAsia along with other two sublineages Ind2001e and Ind2001BD2 were reported in cattle in 2018 from Manikganj ( 27 ). However, the present isolates were collected from different host buffaloes within the same geographical region and year. The farms were not vaccinated against FMD and practiced mixed farming, housing both cattle and buffalo. Recently, the farm experienced an FMD infection among the cattle. Consequently, there was a high probability of FMDV transmission from cattle to buffalo ( 28 ). The lineage Ind2001 has been categorized into four distinct sublineages: Ind2001a, Ind2001b, Ind2001c, Ind2001d, and Ind2001e ( 8 ). Sublineage Ind2001e, is the most prevalent and dominant sublineage in Bangladesh since 2009 ( 9 , 12 , 25 ). Recently, another novel sublineage MYMBD21 under SA-2018 was identified in the sample from 2019 to 2021 in Mymensingh, Bangladesh ( 12 ). Therefore, five distinct lineages of the FMDV serotype O circulating in the Indian subcontinent: SA-2018, Ind2001, PanAsia, PanAsia-2, and Ind2011, with potential cross-border transmission among neighboring countries ( 10 , 12, 25 ). The O/India/R2/75 strain, used in India's national FMD control program as the primary component of the trivalent vaccine, serves as a relevant reference to assess antigenic variation and implications for vaccine efficacy ( 29 ). The VP1 region amino acid sequence of vaccine strain (O/India/R2/75) and BAN 450 isolates differed in 3.75% amino acids and BAN 450 isolates had mutations in both BC loop and C-terminal region. These mutations in the antigenic sites of the BAN 450 isolates may contribute to future vaccine escape by field viruses ( 30 ). As a result, the co-circulation of several sublineages with ongoing mutations that differ from the vaccine strains may pose a significant threat to future outbreaks with high mortality rates. Vaccine failures will have a direct effect on Bangladesh's livestock industry. However, regular surveillance of FMD, along with the study of its genomic and antigenic evolution, and the implementation of an effective national vaccination program, are essential to limit the spread of FMD in Bangladesh. CONCLUSION The emerging sublineage Ind2001e of FMDV serotype O is circulating in buffalo in Manikganj district of Bangladesh. There are several mutations in the VP1 amino acid sequences as compared to vaccination strains, which could make the current FMD situation in Bangladesh even more complicated. The emergence of multiple sublineages of FMDV serotype O in multiple susceptible hosts is a concern for the FMD control program. Emphasis should be placed on increasing vaccination coverage in buffaloes, similar to the vaccination efforts in cattle, by motivating buffalo farmers and continuously monitoring FMD in buffaloes through evaluating the genomic and antigenic variation of FMDV. CONFLICT OF INTEREST
DISCUSSION FMD affects all cloven hoof animal species in Bangladesh every year, especially cattle and buffalo. It has been reported that the genetic evolution of FMDV leads to a consistently increasing tendency of infection and mortality ( 24 , 25 ). Buffalo were perceived as having a lower susceptibility to diseases, and farmers considered buffaloes resistant to FMDV ( 26 ). Evidence indicates that Bangladeshi buffalo farmers occasionally experience outbreaks of FMD, although symptoms are less severe than in cattle ( 6 ). In recent outbreaks, FMDV serotype O has been the predominant serotype in both cattle and buffalo in Bangladesh ( 12 ). However, there is a lack of comprehensive genetic information on FMDV serotype O in buffalo as compared to cattle in Bangladesh. The study aimed to investigate the evolutionary relationships of FMDV serotype O circulating in buffalo in Bangladesh and to compare the amino acid mutation against the vaccine strain based on the VP1 coding sequence of the genome. The VP1 sequencing analysis indicated that the lineage Ind2001 has been predominantly circulating in Bangladeshi cattle since 2009 ( 9 , 25 ). The study identified an emerging novel sublineage Ind2001e (previous name Ind2001BD1) under the ME-SA topotype circulating in both buffalo and cattle. The sublineages Ind2001e and Ind2001BD2 were first identified in samples collected between 2012 and 2016 by Siddique et al. ( 9 ). Later, lineage PanAsia along with other two sublineages Ind2001e and Ind2001BD2 were reported in cattle in 2018 from Manikganj ( 27 ). However, the present isolates were collected from different host buffaloes within the same geographical region and year. The farms were not vaccinated against FMD and practiced mixed farming, housing both cattle and buffalo. Recently, the farm experienced an FMD infection among the cattle. Consequently, there was a high probability of FMDV transmission from cattle to buffalo ( 28 ). The lineage Ind2001 has been categorized into four distinct sublineages: Ind2001a, Ind2001b, Ind2001c, Ind2001d, and Ind2001e ( 8 ). Sublineage Ind2001e, is the most prevalent and dominant sublineage in Bangladesh since 2009 ( 9 , 12 , 25 ). Recently, another novel sublineage MYMBD21 under SA-2018 was identified in the sample from 2019 to 2021 in Mymensingh, Bangladesh ( 12 ). Therefore, five distinct lineages of the FMDV serotype O circulating in the Indian subcontinent: SA-2018, Ind2001, PanAsia, PanAsia-2, and Ind2011, with potential cross-border transmission among neighboring countries ( 10 , 12, 25 ). The O/India/R2/75 strain, used in India's national FMD control program as the primary component of the trivalent vaccine, serves as a relevant reference to assess antigenic variation and implications for vaccine efficacy ( 29 ). The VP1 region amino acid sequence of vaccine strain (O/India/R2/75) and BAN 450 isolates differed in 3.75% amino acids and BAN 450 isolates had mutations in both BC loop and C-terminal region. These mutations in the antigenic sites of the BAN 450 isolates may contribute to future vaccine escape by field viruses ( 30 ). As a result, the co-circulation of several sublineages with ongoing mutations that differ from the vaccine strains may pose a significant threat to future outbreaks with high mortality rates. Vaccine failures will have a direct effect on Bangladesh's livestock industry. However, regular surveillance of FMD, along with the study of its genomic and antigenic evolution, and the implementation of an effective national vaccination program, are essential to limit the spread of FMD in Bangladesh. CONCLUSION The emerging sublineage Ind2001e of FMDV serotype O is circulating in buffalo in Manikganj district of Bangladesh. There are several mutations in the VP1 amino acid sequences as compared to vaccination strains, which could make the current FMD situation in Bangladesh even more complicated. The emergence of multiple sublineages of FMDV serotype O in multiple susceptible hosts is a concern for the FMD control program. Emphasis should be placed on increasing vaccination coverage in buffaloes, similar to the vaccination efforts in cattle, by motivating buffalo farmers and continuously monitoring FMD in buffaloes through evaluating the genomic and antigenic variation of FMDV. CONFLICT OF INTEREST
The authors declare that they have no financial or non-financial conflict of interest regarding authorship and publication of this article. ACKNOWLEDGMENTS
The research was funded by a development project titled "Research on FMD and PPR in Bangladesh" under the Ministry of Fisheries and Livestock, Bangladesh (Award number MOFL-7080). The funding recipient was M. Giasuddin. AUTHORS' CONTRIBUTIONS
MZA made the conceptualization, investigation, methodology, data analysis, and original draft writing. NT and MAT performed the data analysis, writing, reviewing, and editing; MHR and MMM were included in software support, reviewing, and editing. MG contributed to funding acquisition, supervision, methodology, reviewing, and editing. All authors have read and approved the final version of the manuscript.
References
- Kitching, RP (2002). Clinical variation in foot and mouth disease: cattle. Rev Sci Tech. 21(3): 499-502. https://doi.org/10.20506/rst.21.3.1343 PMid:12523690
- Grubman, M.J., Baxt, B. (2004). Foot-and-mouth disease. Clin Microbiol Rev. 17(2): 465-493. https://doi.org/10.1128/CMR.17.2.465-493.2004 PMid:15084510 PMCid:PMC387408
- Knight-Jones, T.J., Rushton, J. (2013). The economic impacts of foot and mouth disease-what are they, how big are they and where do they occur? Prev Vet Med. 112(3-4): 161-173. https://doi.org/10.1016/j.prevetmed.2013.07.013 PMid:23958457 PMCid:PMC3989032
- Bijari, M., Sadeghi-Sefidmazgi, A., Ansari-Lari, M., Ghaffari, M.H. (2025). Productivity, reproduction and economic losses due to foot-and-mouth disease (FMD) in Iranian Holstein cows. Prev Vet Med. 238, 106471. https://doi.org/10.1016/j.prevetmed.2025.106471 PMid: 39978113
- Giasuddin, M., Ali, M.Z., Sayeed, M.A., Islam, E. (2020). Financial loss due to foot and mouth disease outbreak in cattle in some affected areas of Bangladesh. Bangl J Lives Res. 27(1&2): 82-94. https://doi.org/10.3329/bjlr.v27i1.55172
- Nandi, S.P., Rahman, M.Z., Momtaz, S., Sultana, M., Hossain, M.A. (2015). Emergence and distribution of foot‐and‐mouth disease virus serotype A and O in Bangladesh. Transbound Emerg Dis. 62(3): 328-331. https://doi.org/10.1111/tbed.12113 PMid:23734722
- Hemadri,D.,Tosh,C.,Sanyal,A.,Venkataramanan,R. (2002). Emergence of a new strain of type O foot-and-mouth disease virus: its phylogenetic and evolutionary relationship with the PanAsia pandemic strain. Virus Genes. 25(1): 23-34. https://doi.org/10.1023/A:1020165923805 PMid:12206305
- Subramaniam, S., Mohapatra, J.K., Das, B., Sanyal, A., Pattnaik, B. (2015). Genetic and antigenic analysis of foot-and-mouth disease virus serotype O responsible for outbreaks in India during 2013. Infect Genet Evol. 30, 59-64. https://doi.org/10.1016/j.meegid.2014.12.009 PMid:25511252
- Siddique, M.A., Ali, M.R., Alam, A.S.M.R.U., Ullah, H., Rahman, A., Chakrabarty, R.P., Amin, M.A., et al. (2018). Emergence of two novel sublineages Ind2001 BD 1 and Ind2001 BD 2 of foot-and-mouth disease virus serotype O in Bangladesh. Transbound Emerg Dis. 65(4): 1009-1023. https://doi.org/10.1111/tbed.12834 PMid:29457368
- Subramaniam, S., Sanyal, A., Mohapatra, J.K., Sharma, G.K., Biswal, J.K., Ranjan, R., Rout, M., et al. (2013). Emergence of a novel lineage genetically divergent from the predominant Ind2001 lineage of serotype O foot-and-mouth disease virus in India. Infect Genet Evol. 18, 1-7. https://doi.org/10.1016/j.meegid.2013.04.027 PMid:23643555
- Dahiya, S.S., Subramaniam, S., Biswal, J.K., Das, B., Prusty, B.R., Ali, S.Z., Khulape, S.A., et al. (2021). Genetic characterization of foot‐and‐mouth disease virus serotype O isolates collected during 2014-2018 revealed dominance of O/ME‐SA/Ind2001e and the emergence of a novel lineage in India. Transbound Emerg Dis. 68(6): 3498-3508. https://doi.org/10.1111/tbed.13954 PMid:33305514
- Hossain, K.A., Anjume, H., Alam, K.M., Yeamin, A., Akter, S., Hossain, M.A., Sultana, M. (2023). Emergence of a novel sublineage, MYMBD21 under SA-2018 lineage of Foot-and-Mouth Disease Virus serotype O in Bangladesh. Sci Rep. 13(1): 9817. https://doi.org/10.1038/s41598-023-36830-w PMid:37330573 PMCid:PMC10276842
- Hamid, M.A., Siddiky, M.N.A., Rahman, M.A., Hossain, K.M. (2016). Scopes and opportunities of buffalo farming in Bangladesh: A review. SAARC J Agri. 14(2): 63-77. https://doi.org/10.3329/sja.v14i2.31246
- Habib, M.A., Rahman, M.S., Khatun, A., Ali, M.Y. (2023). The scenario of buffalo production and research in Bangladesh. J Buffalo Sci. 12, 28-37. https://doi.org/10.6000/1927-520X.2023.12.04
- Samad, M. (2020). A systematic review of research findings on buffalo health and production published during the last six decades in Bangladesh. J Vet Med OH Res. 2(1): 1-62. https://doi.org/10.36111/jvmohr.2020.2(1).0016
- Vangrysperre, W., De Clercq, K. (1996). Rapid and sensitive polymerase chain reaction based detection and typing of foot-and-mouth disease virus in clinical samples and cell culture isolates, combined with a simultaneous differentiation with other genomically and/or symptomatically related viruses. Arch Virol. 141(2): 331-344. https://doi.org/10.1007/BF01718403 PMid:8634024
- Callens, M., De Clercq, K. (1997). Differentiation of the seven serotypes of foot-and-mouth disease virus by reverse transcriptase polymerase chain reaction. J Virol Methods. 67(1): 35-44. https://doi.org/10.1016/S0166-0934(97)00074-8 PMid:9274816
- Amadori, M., Volpe, G., Defilippi, P., Berneri, C. (1997). Phenotypic features of BHK-21 cells used for production of foot-and-mouth disease vaccine. Biologicals. 25(1): 65-73. https://doi.org/10.1006/biol.1996.006 PMid:9167010
- Knowles, N.J., Wadsworth, J., Bachanek-Bankowska, K., King, D.P. (2016). VP1 sequencing protocol for foot and mouth disease virus molecular epidemiology. Rev Sci Tech. 35(3): 741-755. https://doi.org/10.20506/rst.35.3.2565 PMid:28332654
- Altschul, S.F., Gish, W., Miller, W., Myers, E.W., Lipman, D.J. (1990). Basic local alignment search tool. J Mol Biol. 215(3): 403-410. https://doi.org/10.1016/S0022-2836(05)80360-2 PMid:2231712
- Thompson, J.D., Higgins, D.G., Gibson, T.J. (1994). CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22(22): 4673-4680. https://doi.org/10.1093/nar/22.22.4673 PMid:7984417 PMCid:PMC308517
- Tamura, K., Stecher, G., Kumar, S. (2021). MEGA11: Molecular evolutionary genetics analysis version 11. Mol Biol Evol. 38(7): 3022-3027. https://doi.org/10.1093/molbev/msab120 PMid:33892491 PMCid:PMC8233496
- Tamura, K., Nei, M. (1993). Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 10(3): 512-526.
- WAHID. (2009). OIE World Animal Health Database. c2024 [cited 2024 October 17]. https://wahis.woah.org/#/event-management
- Loth,L.,Osmani,M.G.,Kalam,M.A.,Chakraborty,R.K., Wadsworth, J., Knowles, N.J., Hammond, J.M., Benigno, C. (2011). Molecular characterization of foot‐and‐mouth disease virus: implications for disease control in Bangladesh. Transbound Emerg Dis. 58(3): 240-246. https://doi.org/10.1111/j.1865-1682.2011.01206.x PMid:21320294
- Michel, A.L., Bengis, R.G. (2012). The African buffalo: a villain for inter-species spread of infectious diseases in southern Africa. Onderstepoort J Vet Res. 79(2): 453. https://doi.org/10.4102/ojvr.v79i2.453 PMid:23327373
- Ali, M.Z., Giasuddin, M. (2020). Detection of an emerging novel sublineage Ind2001BD1 and lineage PanAsia of foot-and-mouth disease virus serotype O in cattle in Manikgonj district of Bangladesh, 2018. Open Vet J. 10(3): 347-353. https://doi.org/10.4314/ovj.v10i3.14
- Miguel, E., Grosbois, V., Fritz, H., Caron, A., de Garine‐Wichatitsky, M., Nicod, F., Loveridge, A.J., et al. (2017). Drivers of foot‐and‐mouth disease in cattle at wild/domestic interface: Insights from farmers, buffalo and lions. Divers Distrib. 23(9): 1018-1030. https://doi.org/10.1111/ddi.12585 PMid:32313434 PMCid:PMC7163638
- Upadhyaya, S., Mahapatra, M., Mioulet, V., Parida, S. (2021). Molecular basis of antigenic drift in serotype O foot-and-mouth disease viruses (2013-2018) from Southeast Asia. Viruses. 13(9): 1886. https://doi.org/10.3390/v13091886 PMid:34578467 PMCid:PMC8473337
- Brito, B.P., Rodriguez, L.L., Hammond, J.M., Pinto, J., Perez, A.M. (2017). Review of the global distribution of foot‐and‐mouth disease virus from 2007 to 2014. Transbound Emerg Dis. 64(2): 316-332. https://doi.org/10.1111/tbed.12373 PMid:25996568
Copyright
© 2025 Ali MZ This is an open-access article published under the terms of the Creative Commons Attribution License which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of Interest Statement
The authors declare that they have no financial or non-financial conflict of interest regarding authorship and publication of this article.
Citation Information
Macedonian Veterinary Review. Volume 48, Issue 2, Pages 229-237, e-ISSN 1857-7415, p-ISSN 1409-7621, DOI: https://doi.org/10.2478/macvetrev-2025-0028

 10.2478/macvetrev-2025-0028
10.2478/macvetrev-2025-0028






